Björkling group

Targeted medicinal chemistry towards antibiotics and anti-cancer compounds and Focus on compounds with aim to overcome drug-resistance
-
European Gram Negative Antibacterial Engine (ENABLE)
In ENABLE, http://nd4bb-enable.eu/ we currently work with the optimization of AMPs in close collaboration with external partners. The aim is a focused optimization of activity, toxicology and physical chemical properties to move hit compounds towards lead structures.
In addition, supportive chemistry is performed in other ENABLE programs such as a program aiming for Metallo-β-lactamase inhibitors to be used in combination with carbapenems to overcome emerging drug resistance. -
Center for Peptide based Antibiotics (Cepan)
In Cepan, https://cepan.ku.dk/ current work is focused around the discovery and development of antimicrobial peptides and peptidomimetics with intracellular targets. AMPs with potent antibacterial activity in key Gram-negative bacteria have been discovered. Key properties are currently optimized such as cell permeability, toxicity and stability. The cell penetrating peptides are also used to promote uptake of antisense PNA for inhibition of essential bacterial mRNA thus causing cell death. -
SPRINGBOARD a H2020 Spreading Excellence program
SPRINGBOARD, https://springboard.osi.lv/ aim at exchanging excellence and developing research collaborations between Latvian Institute of Organic Synthesis and University of Copenhagen, University of Antwerp, University of Florence and University of Helsinki within the field of antimicrobial drug research. University of Copenhagen will in a combined effort with the SPRINGBOARD partners provide knowledge in discovery and optimization of antimicrobial peptides and tackle the absolute critical task of antibiotic drug resistance. Research visits to promote these efforts is part of the program.
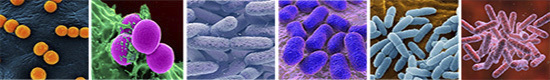
Picture above: ESKAPE pathogens: Enterococcus faecium (1), Staphylococcus aureus (2), Klebsiella pneumoniae (3), Acinetobacter baumannii (4), Pseudomonas aeruginosa (5), Enterobacter cloacae (6)
-
Caught in translation - targeting disease ribosomes
Many diseases are related to defects in ribosomes. These ribosomopathies, although involving ubiquitous fundamental cellular processes, display various clinical manifestations such as short stature, cancer predisposition or hematological disorders. The ribosome is an attractive and well-characterized drug target e.g. many antibiotics exploit minor structural differences to selectively target bacterial (prokaryotic) ribosomes. However, designing compounds capable of altering dysfunctional eukaryotic ribosomes remains a challenge and constitutes a largely unexplored territory. The project is based on the hypothesis that specific pathogenic processes are associated with certain ribosome modifications and the ultimate goal is to discover ligands selectively targeting post-transcriptionally modified ribosomes. This will be explored using a multidisciplinary approach combining computational modelling, medicinal chemistry and biochemistry.
Additional programs
i) Proof of concept study on ‘Translation inhibitors as novel antibiotics’
ii) ‘Structure-based design of Cytochrome P450 17A1 inhibitors’ for the treatment of prostate cancer
iii) ‘Antibody drug conjugates with Nicotinamide phosphoribosyl transferase
(NAMPT) inhibitors as payload’ for treatment of cancer.
Group members
| Name | Title | Phone |
|---|







